Introducing the BioFresh freshwater biodiversity information platform and data portal

Steppe landscape. Image: Creative Commons: Steyr / Picasa
 By Dr. Aaike De Wever, Science Officer for the BioFresh project at the Royal Belgian Institute of Natural Science, and co-ordinator of the freshwater biodiversity data workflow and creation of the public data portal. He can be emailed at data@freshwaterbiodiversity.eu and he tweets @biofreshdata.
By Dr. Aaike De Wever, Science Officer for the BioFresh project at the Royal Belgian Institute of Natural Science, and co-ordinator of the freshwater biodiversity data workflow and creation of the public data portal. He can be emailed at data@freshwaterbiodiversity.eu and he tweets @biofreshdata.
—
When I started to work on the BioFresh project a year ago, I still very much needed to figure out what a freshwater biodiversity information platform and data portal was going to look like. Obviously I immediately had a closer look at the work of my colleagues at the Royal Belgian Institute of Natural Science and the Belgian Biodiversity Platform, especially at SCAR-MarBIN (initiative on marine Antarctic biodiversity data) and the Freshwater Animal Diversity Assessment (FADA; authoritative species lists for freshwater organism groups). After some discussion and reading, however, I soon came to realize that the aim for BioFresh was somewhat different…
The project description reads:
“The freshwater biodiversity information platform is supposed to bring together, and make publicly available, the vast amount of information on freshwater biodiversity currently scattered among a wide range of databases. Previously dispersed and inaccessible data will thus be made available to policy makers, scientists, planners and practitioners. Integration of these data in scientific analyses will lead to better insights in the status and trends of freshwater biodiversity and its ecosystem services and provide scientific support for its management.”
Now, so far the theory, but how do we actually achieve this goal?
A first big task is obviously to identify freshwater biodiversity datasets that are relevant for the scientist within BioFresh and for the scientific community at large. At this stage we are bringing this information together in a metadatabase (a database with information on databases) and are welcoming any information on databases from data holders[1]. This metadatabase aims to make datasets more easily discoverable and aid publication and archiving of datasets. The next step would be to integrate this data in a central data portal.
The first type of data we are integrating in the data portal is primary biodiversity data (taxon-occurrences), which is available through the Global Biodiversity Information Facility. The standards for and the exchange of this type of data through networks of data providers are well established, so there’s no need to reinvent the wheel there. All the more challenging is the integration of other types of data. Up to now we have integrated data from FADA, have made plans to link up with Catalogue of Life, PESI, FishBase and IUCN Red List of Threatened Species, but other types of data such as data from environmental surveys are still under consideration.
A second challenge is making sure we are actually reaching our target audience of scientists, policy maker, planners and practitioners. Given the scientific work within BioFresh and the fact that I am a scientist myself, getting opinions from scientists on what they need and how to make the portal useful for them is relatively easy. Our aim is to assist in data discovery and acquisition, identifying data gaps and stimulate data digitization.
A key requirement for attracting scientists is to ensure good quality data, which is obviously a major challenge and will require us to find the right balance between offering as much data as possible, making sure it is regularly updated and providing detailed data quality information. We still have a long way to go in this respect, but it is certainly high on our agenda.
Reaching out to a wider public of policy maker, planners and practitioners is equally important, but has unfortunately not received a lot of attention up to now. We hope to improve this once we start integrating policy relevant tools and scientific results including the Climate Vulnerability Index and Freshwater Key Biodiversity Areas. In any case we are very much interested to highlight the importance and usefulness of projects like the portal to these groups, looking to start dialogue on how we can effectively translate scientific research into policy making and conservation planning.
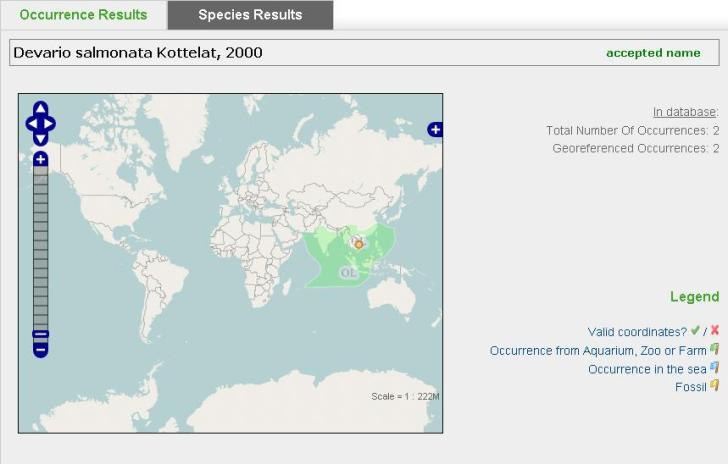
Finally, let’s have a look at the portal itself. During the first year we worked on it, we focused on the basics, making sure we had a solid backbone on which we could further extend and as mentioned above, it is mainly targeted at scientists for now. Visitors can search for information on species belonging to the following organism groups: Halacaridae, Hydroida, Cladocera, Copepoda, Mysidacea, Ephemeroptera, Plecoptera, Macrophytes, Bivalvia, Nematomorpha, Rotifera, Fish and Mammals covering over 32’000 species. For each of these species we offer information on the status of the scientific name and the faunistic region in which the species occurs (based on the information from FADA-experts) and the occurrence data available on GBIF.
Implementation of advanced search capabilities and data download functionality are the major basic features that are still on our to-do list. We recently started planning the integration of models and tools that will be developed within BioFresh and hope to start on this quite soon!
Any feedback on the current state of the portal would be very welcome and will help us to further improve and expand it. You can reach us at data@freshwaterbiodiversity.eu.
[1] If you are holding data that may be relevant for freshwater biodiversity research, we welcome you to contact us at data@freshwaterbiodiversity.eu.
—
Aaike De Wever (PhD, m) is Science Officer for the BioFresh project at the RBINS and will coordinate the data workflow and creation of the public data portal. He is the central contact for data providers and will closely collaborate with the teams at BOKU and WorldFish for technical quality control and digitalization of data. Aaike has a background on aquatic microbial ecology in freshwater environments. For his PhD, he studied the microbial food web in Lake Tanganyika and has experience with challenging experimental field work, molecular ecology and data management. After his PhD, he worked on microbial ecology of Antarctic lakes and teledection of benthic micro-algae on intertidal mudflats.